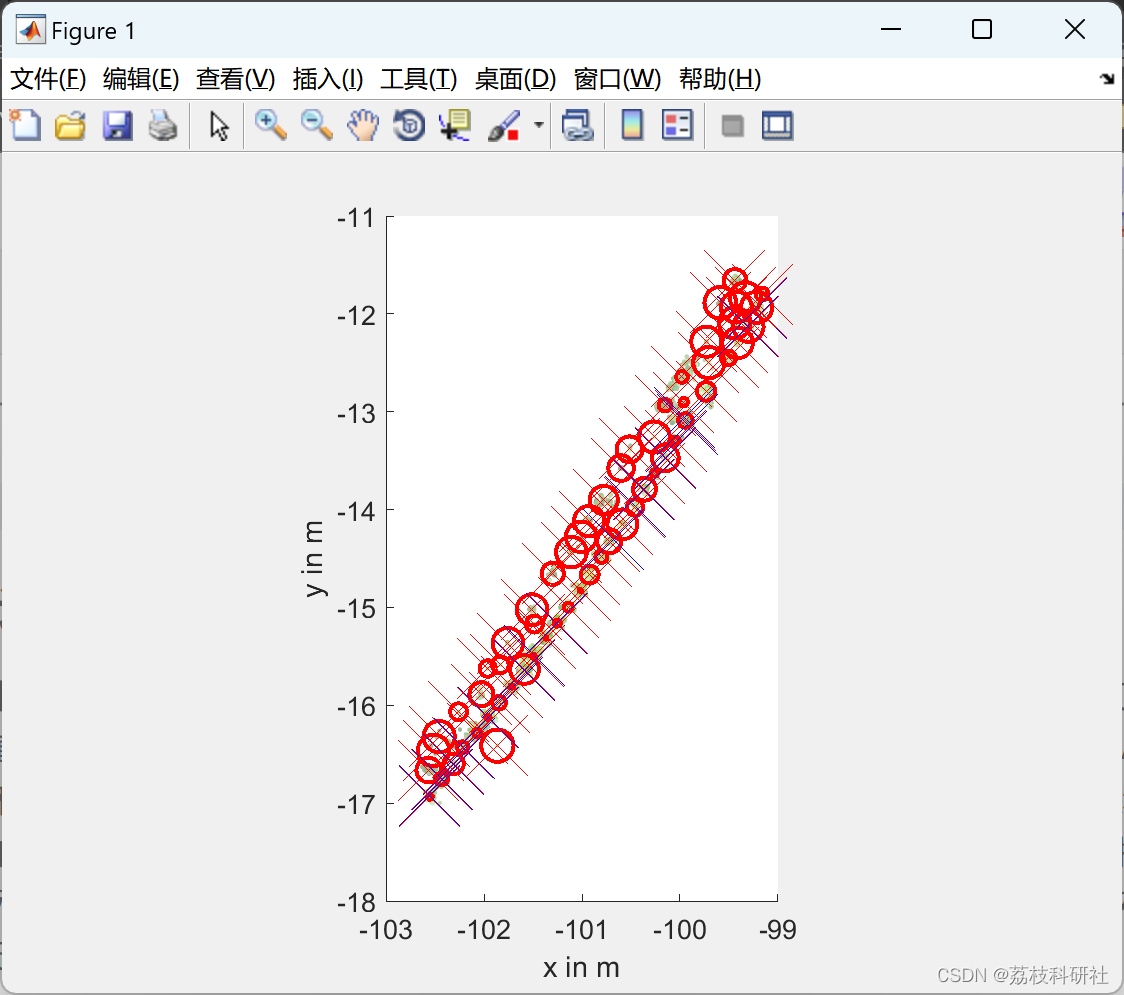
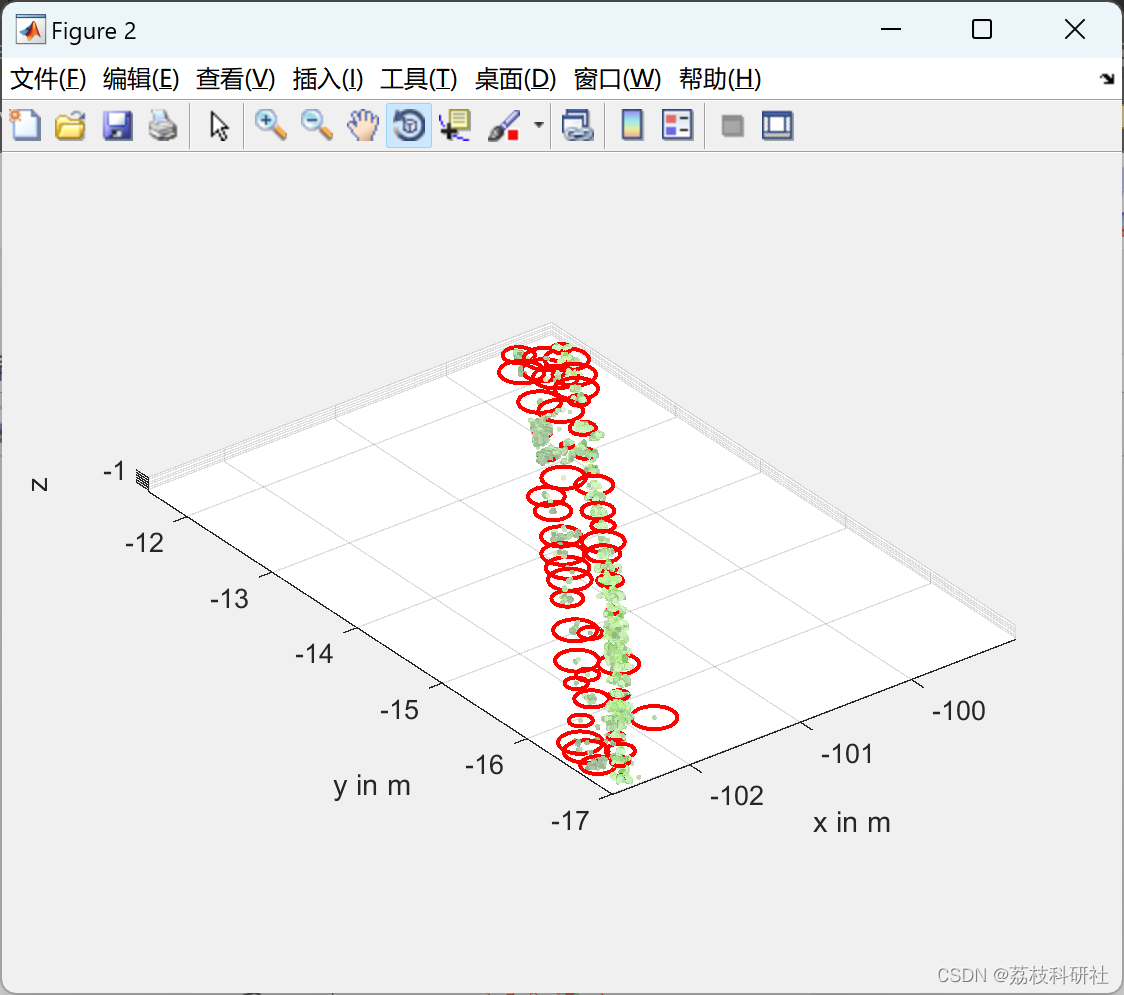
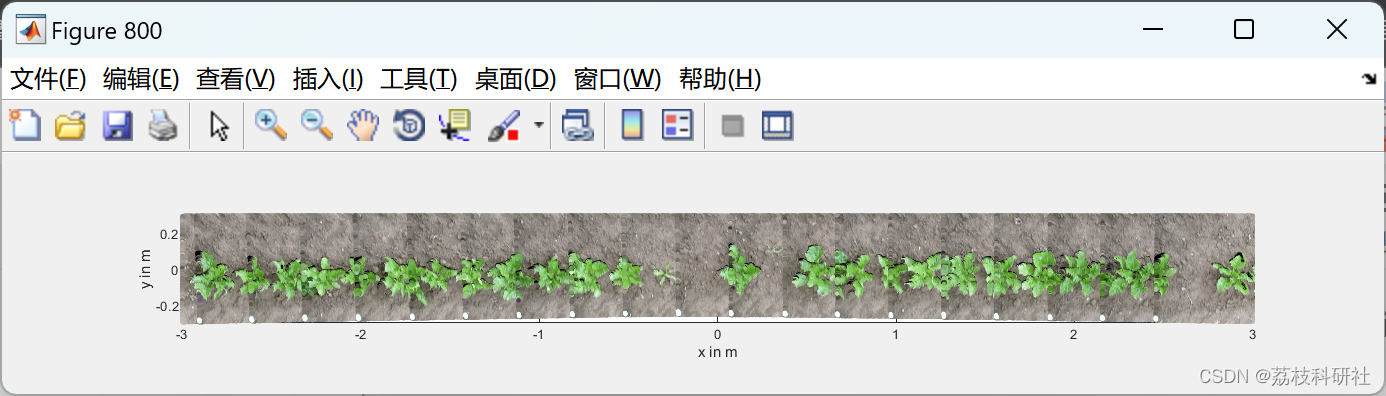
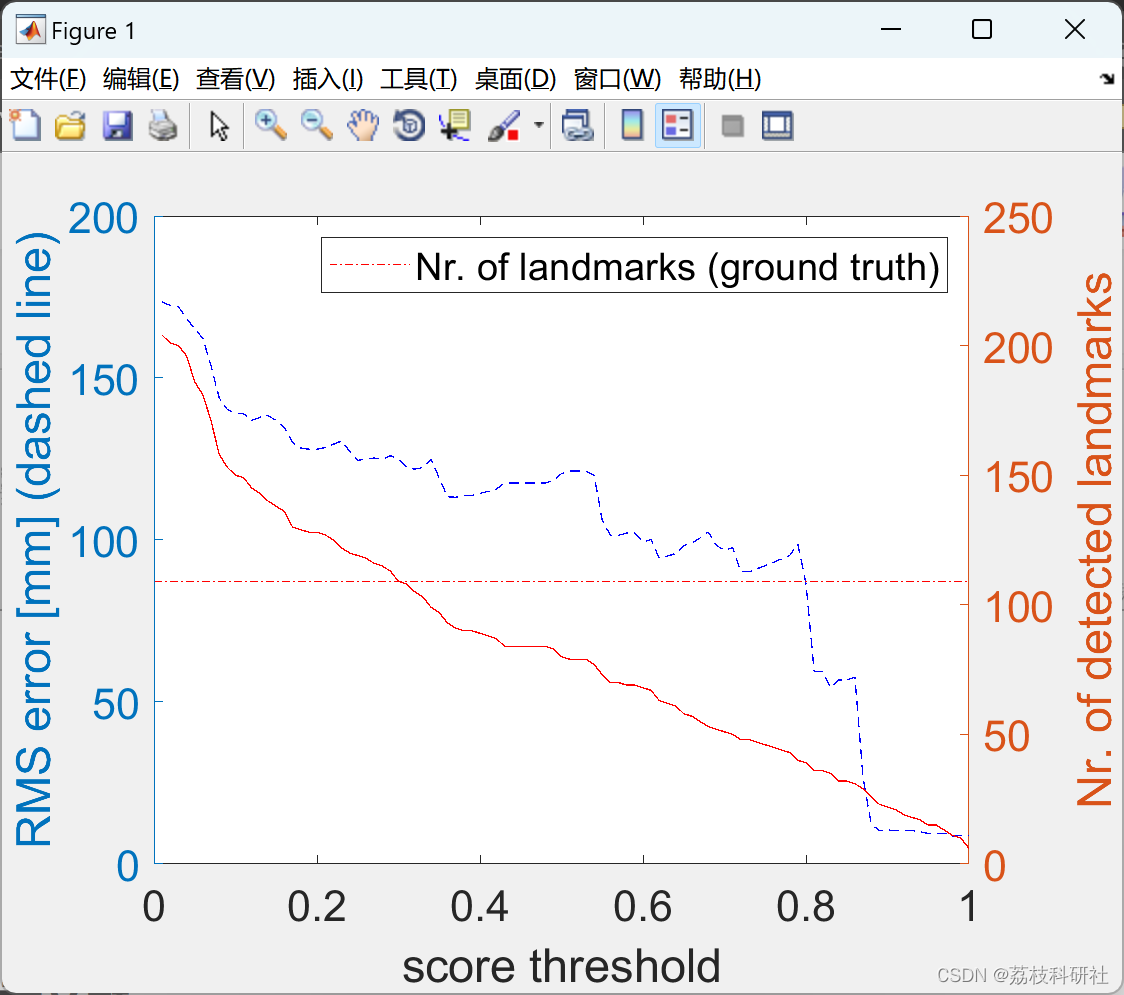
精准农业中使用的 机器人 系统的关键要求之一是能够检测单个植物的确切位置。 我们提出了一种基于从种植园获得的3D点云的新方法,该方法确定确切的植物位置,可用于基于地标的SLAM(同时定位) 和映射)在移动机器人上或用于引导工厂处理系统。 我们方法背后的关键思想是利用同一物种的单个植物之间的几何相似性以及它们播种的规则模式。 因此,我们的方法对于作物以规则模式排列的单一栽培种植园最有用。 在我们的实验研究中,我们表明所提出的方法优于对不同生长阶段记录的真实世界数据进行的传统平均偏移分析。


部分代码:
%% plot examples from all days
for day = 1:nDays
h = figure(day+798);
datasets = days(day).datasets;
data = load(strcat(basepathData, datasets{2}));
pNorm = geometry.normalizePointCloud(data.mapFull);
plot.pc(pNorm(:,1:2), data.mapColorFull, 'markersize', 20);
ax = gca;
ax.FontSize = 5;
ax.XLim = [-3 3];
ax.YLim = [-0.3 0.3];
set(gcf, 'Position', [200.6040 713.8713 691.0099 120.7129])
print(h, '-dpng', [basepathImages 'example_day' num2str(day) '.png'], '-r300')
end
%% do the evaluation for each error_threshold
for threshold_error = [0.01 0.02 0.04]
%%
for day = 1:nDays
%% evaluate each day individually
datasets = days(day).datasets;
% combine individual datasets for each day
dayCombined.landmarksMeanshift = [];
dayCombined.landmarksMeanshiftScores = [];
dayCombined.landmarksApproach = [];
dayCombined.landmarkScores = [];
dayCombined.landmarksManual = [];
dayCombined.map = [];
dayCombined.mapColor = [];
for j = 1:numel(datasets)
datasetPath = [basepathData datasets{j}];
data = load(datasetPath);
%% plant soil segmentation
if ~isfield(data, 'map')
[segmentation, extractorPolygon1, extractorPolygon2] = tools.greenextract(data.mapFull, data.mapColorFull);
data.map = data.mapFull(segmentation,:);
data.mapColor = data.mapColorFull(segmentation,:);
save(datasetPath, '-struct', 'data', 'map', 'mapColor','-append')
end
%% manual landmarks
if ~isfield(data, 'landmarksManual')
h = figure(733);
plot.pc(data.map, data.mapColor, 'markerSize', 20)
datacursormode on
dcm_obj = datacursormode(gcf);
waitfor(msgbox('Click ok when finished labeling plants!'));
cursor_info = getCursorInfo(dcm_obj);
data.landmarksManual = vertcat(cursor_info.Position);
if size (data.landmarksManual,2) == 3
data.landmarksManual = data.landmarksManual(:,1:2);
end
save(datasetPath, '-struct', 'data', 'landmarksManual','-append')
end
if updateLandmarksManual
figure(733);
h = plot.pc(data.map(:,1:2), data.mapColor, 'markerSize', 20);
datacursormode on
dcm_obj = datacursormode(gcf);
hTarget = handle(h);
xdata = get (hTarget,'XData');
ydata = get(hTarget,'YData');
idx = knnsearch([xdata' ydata'], data.landmarksManual);
for tipIdx = 1:size(data.landmarksManual, 1)
hDatatip = dcm_obj.createDatatip(hTarget);
propPointDataCursor = get(hDatatip,'Cursor');
% Move the datatip
propPointDataCursor.DataIndex = idx(tipIdx);
pos = [xdata(idx(tipIdx)) ydata(idx(tipIdx))];
propPointDataCursor.Position = pos;
set(hDatatip,'Position',pos)
end
waitfor(msgbox('Click ok when finished labeling plants!'));
cursor_info = getCursorInfo(dcm_obj);
data.landmarksManual = vertcat(cursor_info.Position);
if size(data.landmarksManual,2) == 3
data.landmarksManual = data.landmarksManual(:,1:2);
end
save(datasetPath, '-struct', 'data', 'landmarksManual','-append')
end
%% tf density approach on whole map
if recalculateApproach && isfield(data, 'landmarksApproach')
data = rmfield(data, 'landmarksApproach');
end
if ~isfield(data, 'landmarksApproach')
[data.landmarksApproach, data.landmarkScores] = optimization.detectPlants(data.map, days(day).args{:});
save(datasetPath, '-struct', 'data', 'landmarksApproach', 'landmarkScores', '-append')
end
%% mean shift baseline
if recalculateMeanshift && isfield(data, 'landmarksMeanshift')
data = rmfield(data, 'landmarksMeanshift');
end
if ~isfield(data, 'landmarksMeanshift')
[data.landmarksMeanshift,~,clusterSizes] = clustering.meanshift(data.map', days(day).args{12});
data.landmarksMeanshiftScores = vector.normalize(cellfun('length',clusterSizes));
save(datasetPath, '-struct', 'data', 'landmarksMeanshift', 'landmarksMeanshiftScores', '-append')
end
%% combine
dayCombined.landmarksMeanshift = [dayCombined.landmarksMeanshift; data.landmarksMeanshift];
dayCombined.landmarksMeanshiftScores = [dayCombined.landmarksMeanshiftScores; data.landmarksMeanshiftScores];
dayCombined.landmarksApproach = [dayCombined.landmarksApproach; data.landmarksApproach];
dayCombined.landmarkScores = [dayCombined.landmarkScores; data.landmarkScores];
dayCombined.landmarksManual = [dayCombined.landmarksManual; data.landmarksManual];
dayCombined.map = [dayCombined.map; data.map];
dayCombined.mapColor = [dayCombined.mapColor; data.mapColor];
end
landmarksMeanshift = dayCombined.landmarksMeanshift;
landmarksMeanshiftScores = dayCombined.landmarksMeanshiftScores;
landmarksApproach = dayCombined.landmarksApproach;
landmarkScores = dayCombined.landmarkScores;
landmarksManual = dayCombined.landmarksManual;
fprintf('day %d: %d labels\n', day,size(landmarksManual,1))
%% calculate errors
[errorMeanshiftVecidx, errorMeanshift] = knnsearch(landmarksManual, landmarksMeanshift(:,1:2));
[errorApproachVecidx, errorApproach] = knnsearch(landmarksManual, landmarksApproach(:,1:2));
errorMeanshiftVec = landmarksManual(errorMeanshiftVecidx,:) - landmarksMeanshift(:,1:2);
errorApproachVec = landmarksManual(errorApproachVecidx,:) - landmarksApproach(:,1:2);
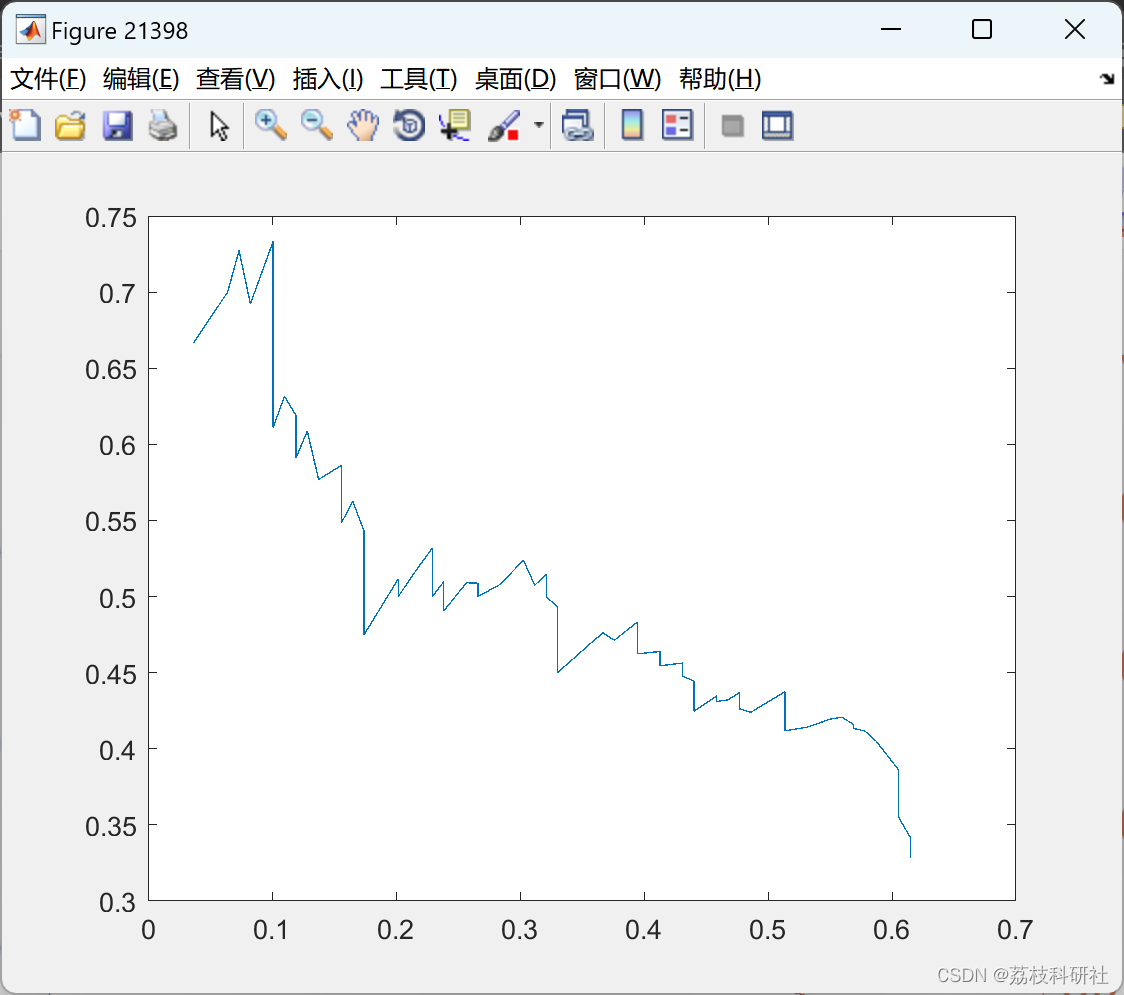
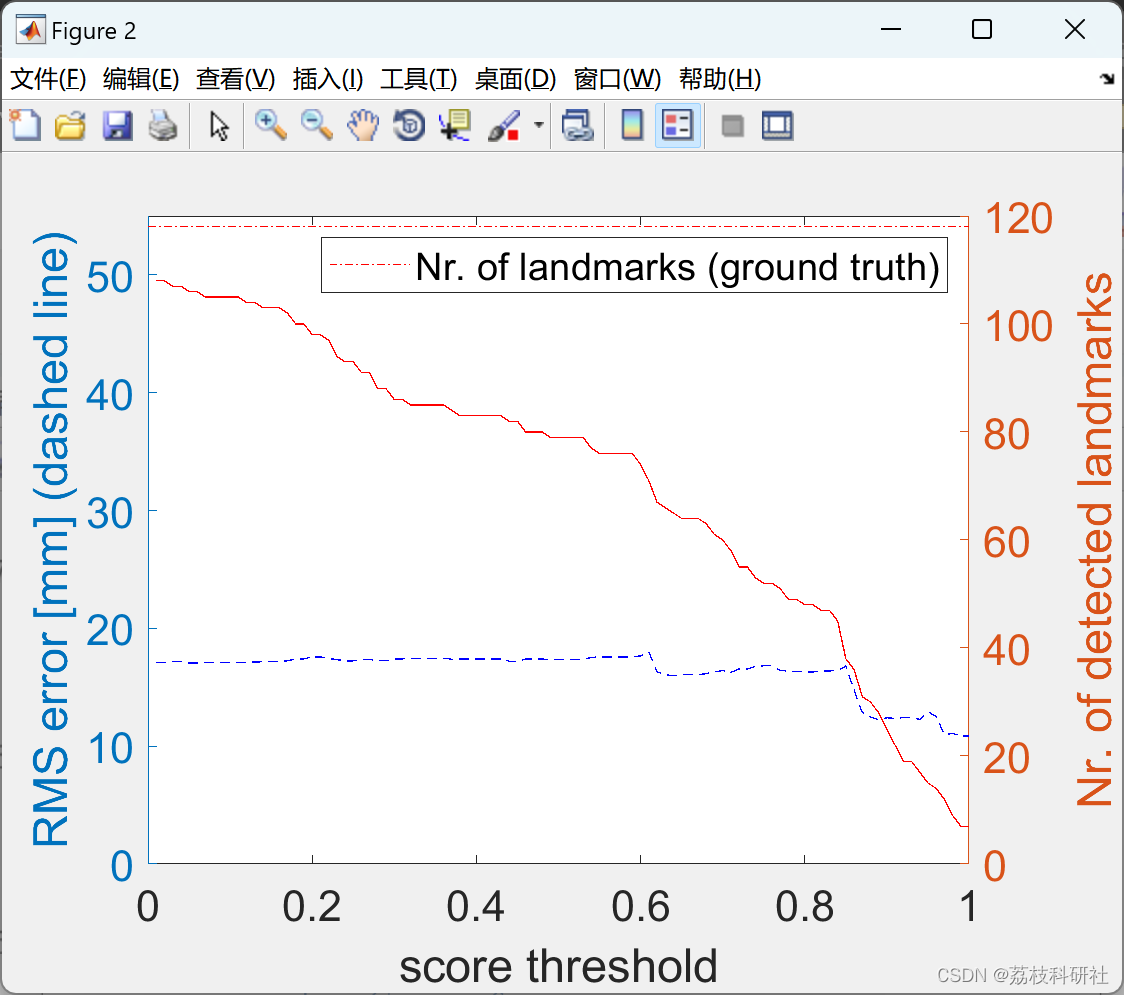
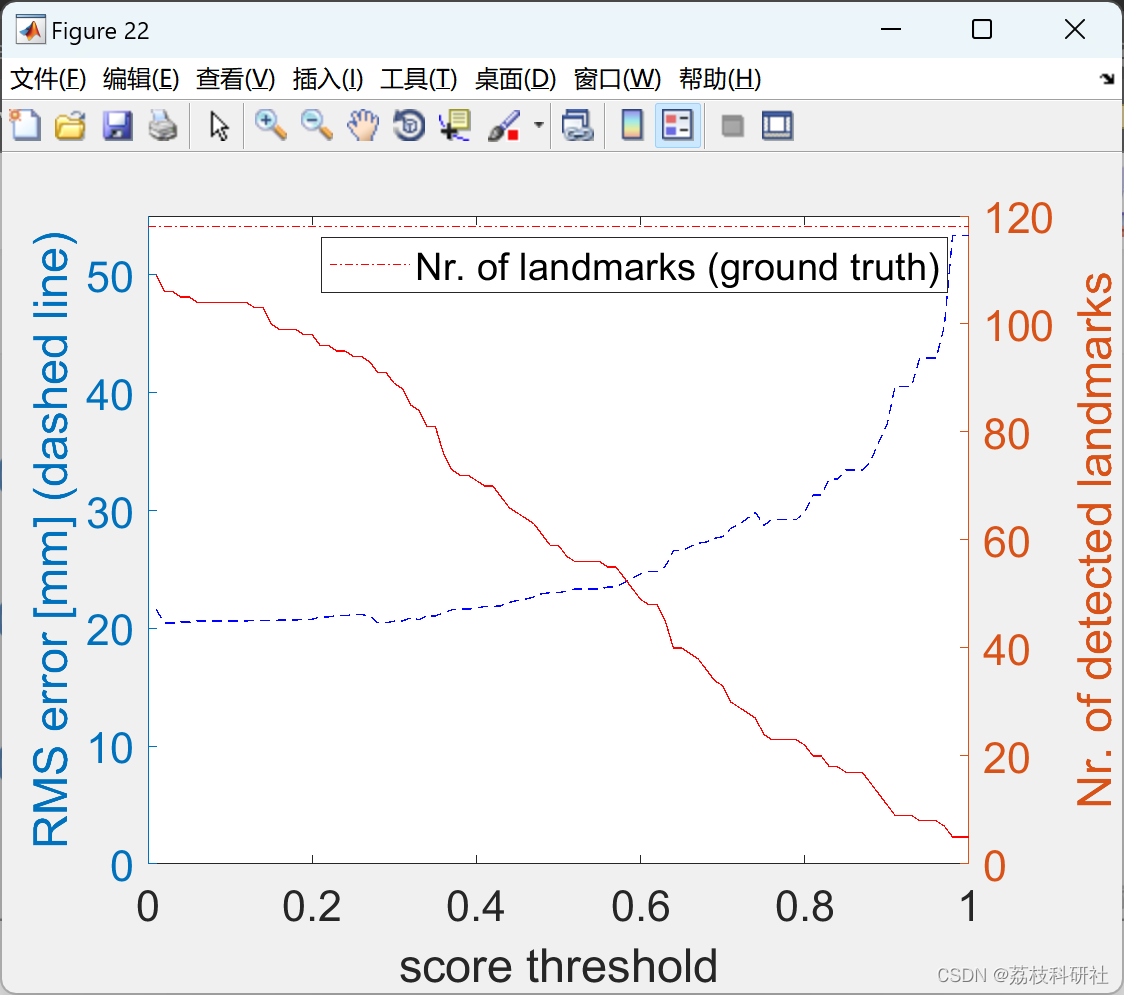
%% eval approach
figure(day); view(2)
thresholds = [];
rms = [];
sizes = [];
for step = 1:100
landmarksApproachSel = landmarksApproach(landmarkScores > ((step-1)/100),:);
[errorApproachVecidx, errorApproach] = knnsearch(landmarksManual, landmarksApproachSel(:,1:2));
sizes(step) = size(landmarksApproachSel,1);
thresholds(step) = step/100;
rms(step) = sqrt(mean(errorApproach.^2));
if thresholds(step) == 0.9
fprintf('day %d, score threshold 0.9, rms %f, detection ratio %f\n', day, rms(step), sizes(step)/size(landmarksManual,1))
end
end
yyaxis left
plot(thresholds,1000*rms, '--b')
xlabel('score threshold')
部分理论来源于网络,如有侵权请联系删除。
[1]Janosch Dobler, Alexander Schaefer, Wolfram Burgard (2018) Symmetry-Based Plant Detection in 3D Point Clouds from Crop Fields
免责声明:本文系网络转载或改编,未找到原创作者,版权归原作者所有。如涉及版权,请联系删